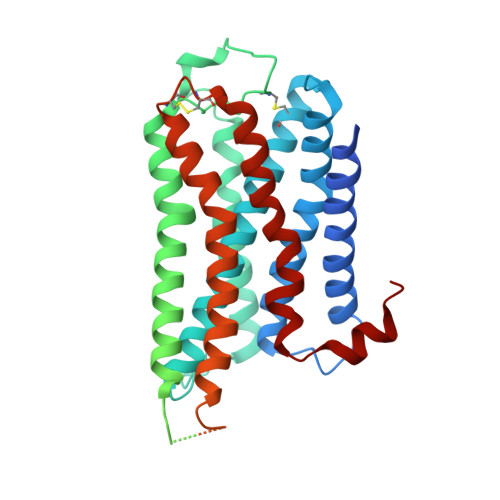
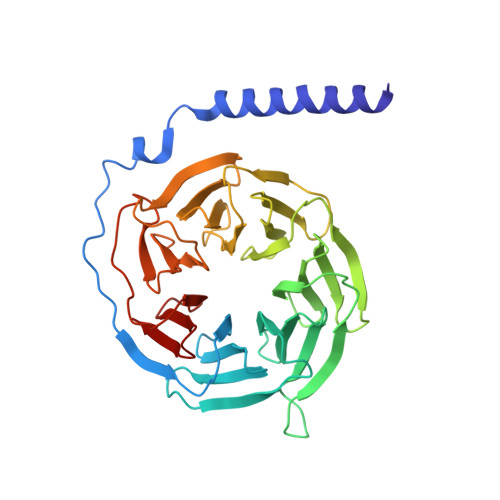
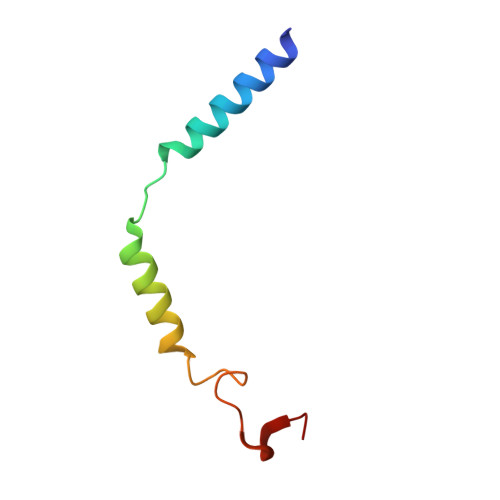
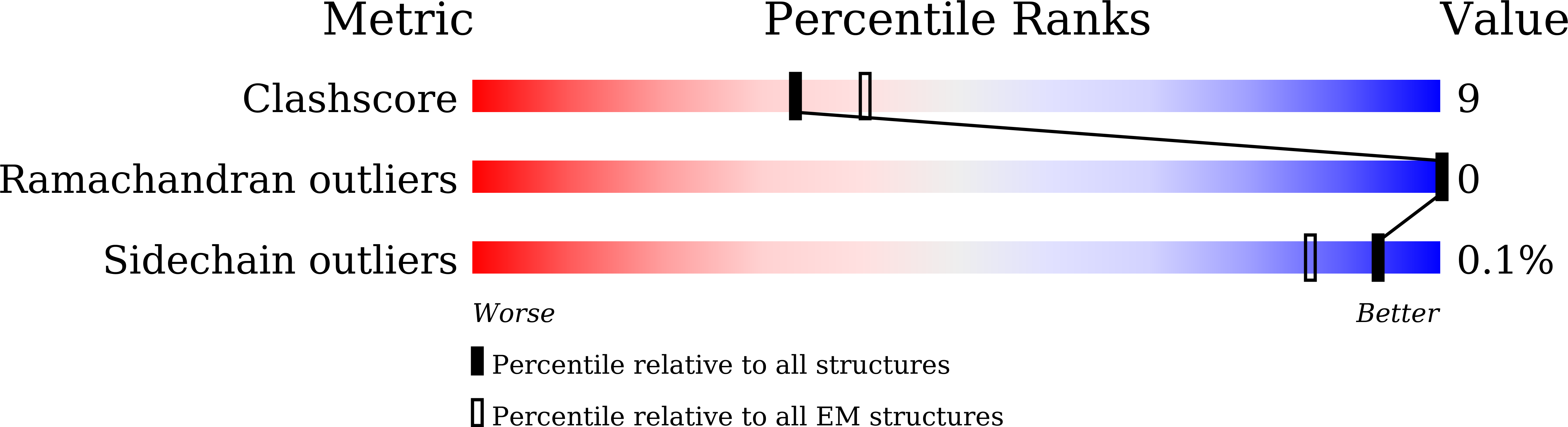Cryo-EM structure of the human histamine H 1 receptor/G q complex.
Xia, R., Wang, N., Xu, Z., Lu, Y., Song, J., Zhang, A., Guo, C., He, Y.(2021) Nat Commun 12: 2086-2086
- PubMed: 33828102
- DOI: https://doi.org/10.1038/s41467-021-22427-2
- Primary Citation of Related Structures:
7DFL - PubMed Abstract:
Histamine receptors play important roles in various pathophysiological conditions and are effective targets for anti-allergy treatment, however the mechanism of receptor activation remain elusive. Here, we present the cryo-electron microscopy (cryo-EM) structure of the human H 1 R in complex with a G q protein in an active conformation via a NanoBiT tethering strategy. The structure reveals that histamine activates receptor via interacting with the key residues of both transmembrane domain 3 (TM3) and TM6 to squash the binding pocket on the extracellular side and to open the cavity on the intracellular side for G q engagement in a model of "squash to activate and expand to deactivate". The structure also reveals features for G q coupling, including the interaction between intracellular loop 2 (ICL2) and the αN-β junction of G q/11 protein. The detailed analysis of our structure will provide a framework for understanding G-protein coupling selectivity and clues for designing novel antihistamines.
Organizational Affiliation:
Laboratory of Receptor Structure and Signaling, The HIT Center for Life Sciences, Harbin Institute of Technology, Harbin, China.